News
Follow us on BlueSky for more News: @bornberglab
23.02.2026
New paper! Orthologs of an essential orphan gene vary in their capacities for function and subcellular localization in Drosophila melanogaster
Prajal H Patel , Lars A. Eicholt , Andreas Lange , Kerry L McDermott , Erich Bornberg-Bauer , Geoffrey D Findlay Molecular Biology and Evolution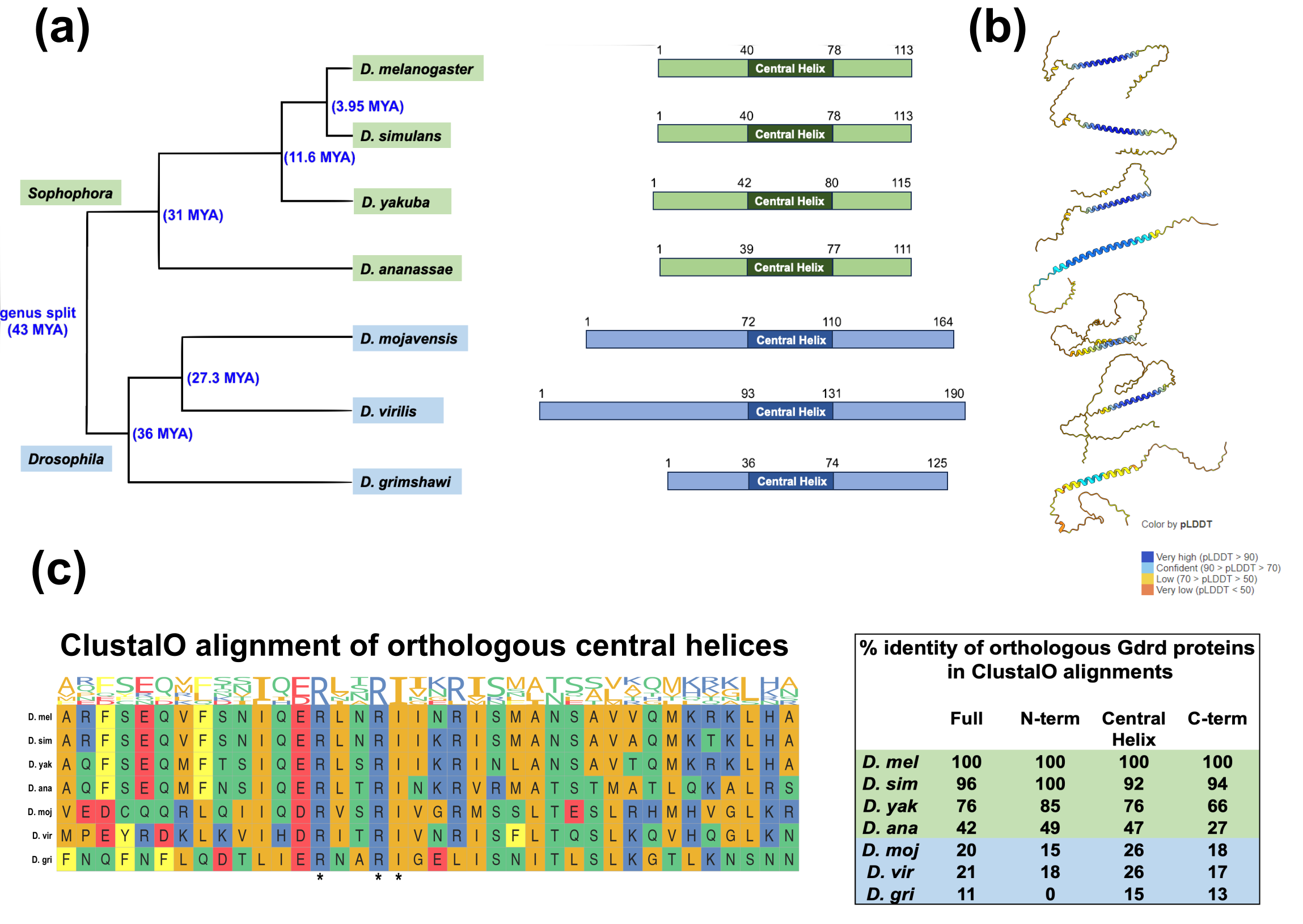
Orphan genes evolve rapidly and often differ strongly between closely related species, making it unclear whether their functions remain conserved over evolutionary time. In a new study, we investigated goddard (gdrd), an orphan gene required for male fertility in Drosophila melanogaster together with our collaborator Geoff Findlay, who led this project. By swapping gdrd orthologs from different Drosophila species into D. melanogaster testes, we found that some highly divergent versions can still rescue fertility, while others, including some from more closely related species, cannot. This suggests that Gdrd acts in a conserved spermatogenesis pathway, but that individual orthologs have undergone lineage-specific functional changes. Cell biological analyses showed that divergent Gdrd proteins still interact with key sperm structures, including axonemes and ring centrioles, although non-rescuing variants often bind more weakly or localize differently. Computational analyses and molecular dynamics simulations further suggest that differences in disordered regions and structural stability help explain why some orthologs retain function while others do not. Overall, the study shows that the ancestral structure and cellular interactions of Gdrd are conserved, but that rapid sequence evolution has led to functional divergence among several species-specific orthologs. You can read the paper here!
08.04.2026
Group Retreat in Diever
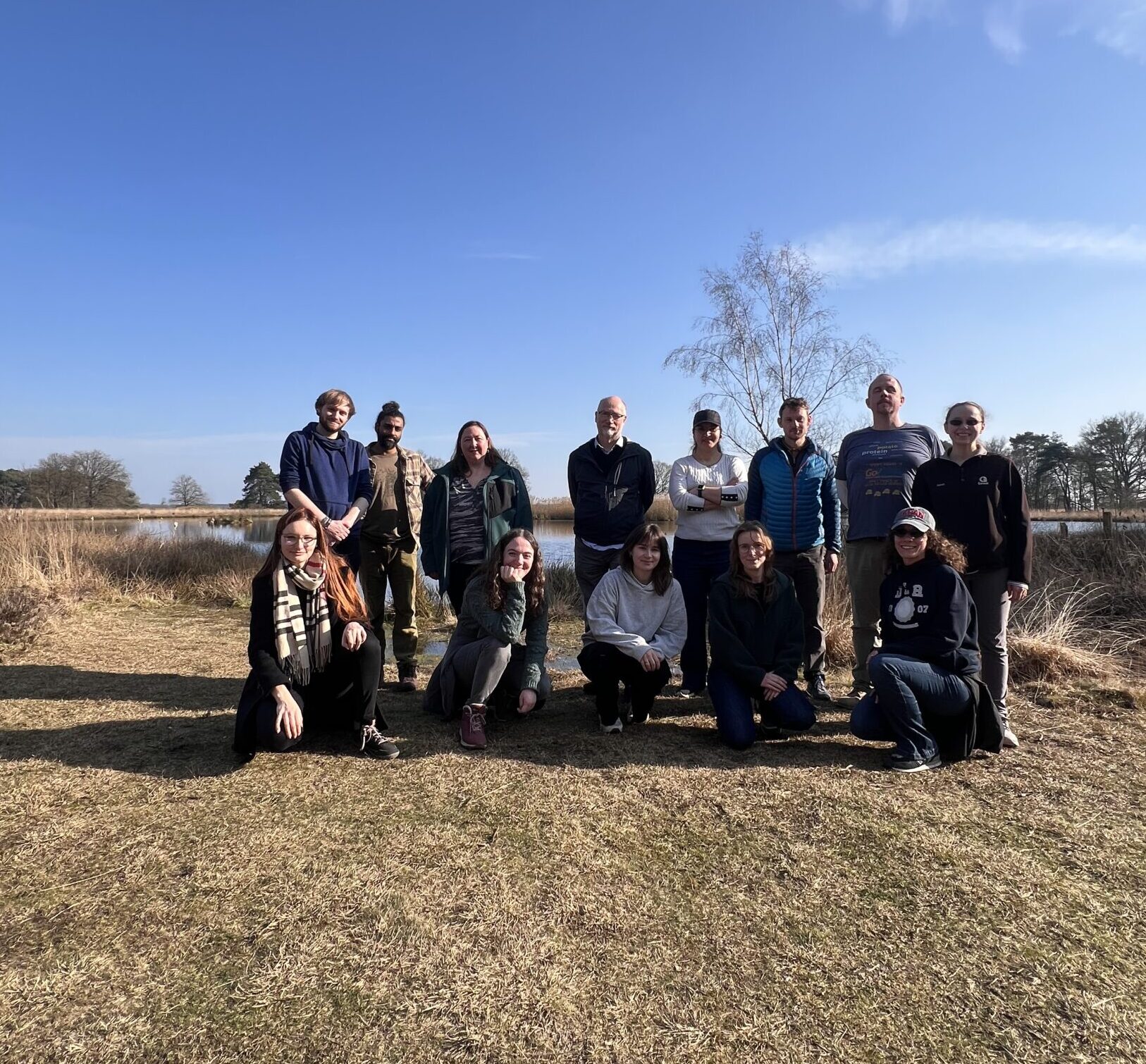

This year, our group traveled to Diever, in the Netherlands. Alongside a series of engaging talks focused on selection, we enjoyed a variety of fun activities, including an escape room and axe throwing. We also took advantage of the sunny weather by going on hikes through the beautiful nature surrounding Diever. You can find more photos here soon.
08.04.2026
DZG Graduate Meeting in Hamburg

Marie traveled to beautiful Hamburg to attend this year’s graduate meeting of the Evolutionary Biology Section of the DZG, where she presented her research about “Comparative Analysis of De Novo Transcript Turnover in Six Drosophila Species.” Many thanks to the organizers for putting together such a nice meeting, featuring great talks and networking opportunities.
23.02.2026
New paper! Studying the evolutionary potential of ancestral aryl sulfatases in the alkaline phosphatase family with droplet microfluidics
Bernard D. G. Eenink , Josephin M. Holstein, Magdalena Heberlein, Carina Dilkaute, Joachim Jose, Florian Hollfelder, Bert van Loo, Erich Bornberg-Bauer , Tomasz S. Kaminski and Andreas Lange Analyst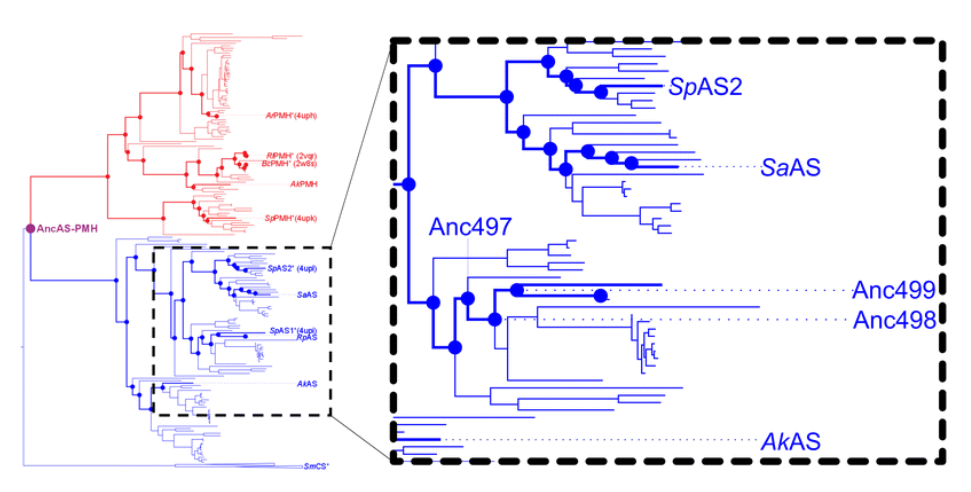
In this paper from Berndjan , they used droplet microfluidics to study the evolutionary potential of aryl sulfatases (a subfamily of the evolutionary old alkaline phosphatase superfamily) and their computationally reconstructed ancestors. They show higher mutation rates gave a wider distribution of active variants but fewer improved variants overall. Moreover they showed differences in the impact of mutation rate, where some enzymes benefited from a higher and others from a lower rate. This highlights the need to test diverse mutagenesis regimes. You can read the paper here!
09.02.2026
New Paper in GBE! Cryptocercus genomes expand knowledge of adaptations to xylophagy and termite sociality
Alun R C Jones , Alina A. Mikhailova , Cédric Aumont, Juliette Berger, Cong Liu, Shulin He, Zongqing Wang, Sylke Winkler, Erich Bornberg-Bauer , Frédéric Legendre, Dino P McMahon, Mark C. Harrison Genome Biology and Evolution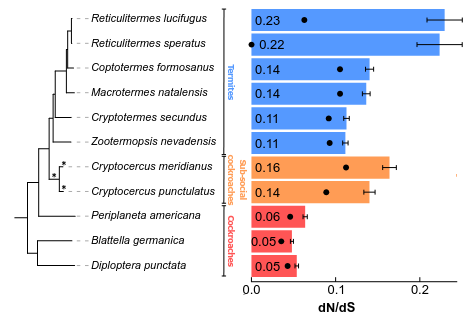
This paper analyzes two high-quality genomes of Cryptocercus, the sister group to all termites, to investigate the transition to subsociality and wood-feeding (xylophagy) in Blattodea. The authors find evidence of relaxed selection in both Cryptocercus and termites, which may indicate a reduction in effective population size in their subsocial ancestor. They also report a reduction in the number of ionotropic receptors and a shift from uni- to bimodal DNA methylation patterns in protein-coding genes in the common ancestor of Cryptocercus and termites. This was previously assumed to have evolved in termites with the emergence of eusociality. Check out the paper here!
28.01.2026
Paper Alert! Emergence and evolution of protein-coding de novo genes
Erich Bornberg-Bauer & Lars A. Eicholt, Nature Reviews Genetics
A new review by Erich and Lars summarizes current understanding of protein-coding de novo genes—genes that originate from previously non-coding DNA. Once considered rare or unlikely, de novo gene birth has become a rapidly growing research area, driven by expanding genome resources, improved comparative genomics, and new functional and structural assays. At the same time, key aspects remain debated: estimates of how often de novo genes arise and persist differ between studies, and their properties can challenge classical expectations about how new genes and proteins emerge. The review discusses how de novo genes are identified (including phylogenetic and synteny-based strategies), which genomic and regulatory features facilitate their emergence, and how population-level gain–loss dynamics shape their fate. It also surveys what is known about the length, intrinsic disorder, structural properties, and cellular functions of proteins encoded by de novo genes, highlighting both experimental evidence and computational advances (including modern structure prediction and large-scale screening approaches). Finally, the article outlines open questions and future directions—aiming to help reconcile differing perspectives and to clarify when, how, and why novel coding sequences become integrated into biological systems over evolutionary time. Read it here!
17.12.2025
Paper Alert! Unravelling the evolution of wood-feeding in termites with 47 high-resolution genome assemblies
Cong Liu, Cédric Aumont, Alina A. Mikhailova , Tracy Audisio, Simon Hellemans, Yi-Ming Weng, Shulin He, Crystal Clitheroe, Zongqing Wang, Ives Haifig, David Sillam-Dussès, Aleš Buček, Gaku Tokuda, Jan Šobotník, , Mark C. Harrison , Dino P. McMahon & Thomas Bourguignon Nature Communications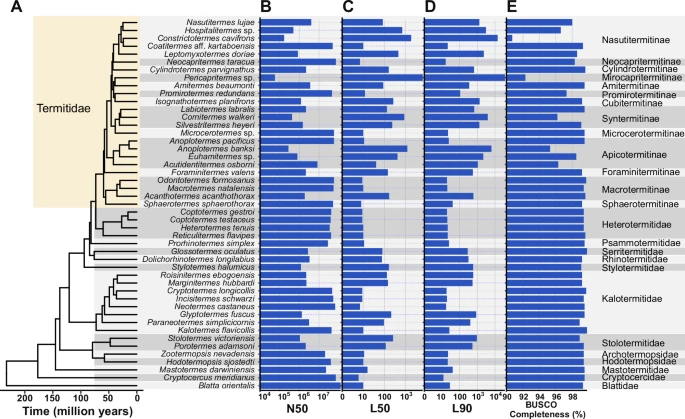
Congratulations to Alina and Mark on this exciting new paper! In this study, they generated 47 new high-quality genomes from termites (45 genomes) and cockroaches (2 genomes), increasing the number of available genomes in this group sixfold and thus filling a major gap in existing data. This dataset allowed the authors to take a closer look at termite genome evolution. They found that members of the Termitidae displayed larger genomes, a higher transposable element content, and higher gene numbers compared to other termites. Notably, this includes an expansion of CAZymes which are the genes involved in lignocellulose degradation. These genomic changes coincide with the origin of soil feeding in Termitidae.
12.12.2025
PhD defense Berndjan Eenink and Alina Mikhailova

Berndjan and Alina both successfully defended their PhD theses this week. Berndjan worked on “Probing the evolvability of ancestral and extant sulfatases using fluorescence-activated droplet sorting (FADS)”. Alina studied the “Genomic basis of sociality in order Blattodea”. Congrats to both of them!
12.12.2025
20 years of IEB: Anniversary Symposium and Christmas Party

We celebrated the 20 year anniversary of our institute with a symposium featuring talks from many alumni. This included several alumni from our group, namely January Weiner, Mark Harrison, Ann Kathrin Huylmans, Bertrand Fouks, and Sonja Grath. The symposium concluded on a festive note with our annual IEB Christmas party.
12.12.2025
Bornberglab Christmas Party 2025

We had our annual Christmas party this week. As usual we started with a movie in the cinema: this year we watched: “Tokyo Godfathers”. After a short stroll over the Christmas market we went for dinner with the whole group as well as some of our alumni.
12.12.2025
Final Project Meeting "Ghost in the protein" in Uppsala

Last week, we wrapped up our Volkswagen Foundation–funded project with a wonderful final meeting in Uppsala, hosted by Ylva Ivarsson’s group. The photo captures the project members from our lab together with colleagues from the groups of Ylva Ivarsson, Klara Hlouchova, and Florian Hollfelder. Shown are: front row (sitting, left to right) Bharat, Isabella, Lars, Tereza, Klara, Theodora; back row (standing, left to right) Jamil, Ylva, Florian, Erich, Filip
04.11.2025
New Paper out! DeNoFo: a file format and toolkit for standardized, comparable de novo gene annotation
Elias Dohmen, Margaux Aubel, Lars A Eicholt, Paul Roginski, Victor Luria, Amir Karger, Anna Grandchamp Bioinformatics
In this study, led by our former postdocs Elias Dohmen and Anna Grandchamp, the authors present a new format designed to standardize the annotation of methods for detecting de novo genes. Complementing this, they developed the DeNoFo toolkit that enables dataset annotation and comparison of the underlying methods across studies. The toolkit is available on GitHub. The study is accompanied by a review that summarizes commonly used de novo gene detection methods and their limitations.
05.09.2025
Annual GEvol Meeting in Münster

Last week, the third annual meeting of the DFG Priority Program “Genomic Basis of Evolutionary Innovations” (GEvol , SPP 2349) was held at the castle in Münster. This meeting marked the end of the first three year funding period of the project which started with a kickoff meeting in Münster in 2022. The first day of the meeting was focused on the renewal, during which all current and prospective GEvol members defended their proposals for the second funding phase. It was a pleasure to welcome Dr. Breitkreuz from the DFG and all reviewers. The achievements of the program’s first phase were met with great appreciation by the reviewers, and the next phase is now eagerly anticipated. The second day of the meeting was dedicated to the projects current PhD students who presented the progress of their projects. Finally, the meeting concluded with the “Long Night of Insects ” at the LWL Museum of Natural History. Attendees could participate in a variety of activities, including a science slam with two GEvol PhD students presenting, an ant farm, educational booths, and a cockroach race.
25.08.2025
ESEB 2025 in Barcelona

Erich and Marie represented the group at ESEB 2025 in Barcelona. Erich gave one talk in the Symposium on “Gene Content Across Genomes: Models and Genomic Data” and another in the Symposium “The future meets the beginning: Synthetic biology, evolution, and the origin of life”. Marie presented her work in the Symposium “Genomic basis of evolutionary innovations” organised by GEvol.
22.07.2025
Brendas Research Module the Earth-Life Science Institute (ELSI) in Japan

We are proud of our master student Brenda, who completed a research module at the Earth-Life Science Institute (ELSI) in Japan. She worked on enzymatic nucleotide phosphorylation using polyphosphates as phosphate donors, focusing on the PPK2 Class III enzyme MAN. The project aimed to better understand ancient enzymes that may have played a key role in the origin of life by catalyzing the conversion of NMPs to NTPs. Now she is off to new adventures, doing her master thesis at the German Aerospace Center (DLR), where she will analyze proteomics and transcriptomics data from neural cells exposed to simulated microgravity or hypergravity. Her goal is to develop a bioinformatics pipeline to interpret “omics” datasets and identify key molecular changes in astrocytes, motor neurons, and brain organoids. We wish her great success on her next steps!
15.04.2025
New Paper! Expression of De Novo Open Reading Frames in Natural Populations of Drosophila melanogaster
Amanda Glaser-Schmitt, Marie Lebherz, Ezgi Saydam, Erich Bornberg-Bauer, John Parsch Journal of Experimental Zoology Part B: Molecular and Developmental Evolution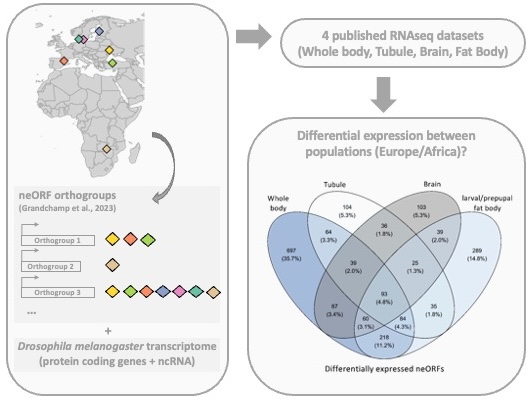
Together with the Parsch lab , we studied the expression of neORFs (newly evolved expressed ORFs) in Drosophila populations, in comparison to protein-coding genes and ncRNA. We used publicly available RNA-seq data, including whole-bodies, two adult tissues, and three larval/developmental stages from a single tissue. We found that about half of the previously detected neORFs were expressed in each dataset and show greater expression divergence between populations than protein-coding genes or ncRNA. In contrast, we observe less sex-biased expression in neORFs compared to the other two gene classes. Notably, the highest number of sex biased neORFs was detected in the whole-body samples, which might be explained by the presence of the gonads. Finally, we found that neORFs shared between multiple lines in the original dataset were more likely to be (differentially) expressed in the four studied datasets, suggesting that they are more likely to be functional.
25.03.2024
3rd Münster Evolution Meeting (MEM)

This year the 3rd Münster Evolution Meeting took place in the Schloss. Like in the last years the conference covered a broad range of topics ranging from talks about sex chromosome evolution to speciation in bumblebees. From our group Marie Lebherz, Bharat Ravi and Lars Eicholt gave talks. Elias Dohmen, Carsten Kemena, Sarah Lucas and Margaux Aubel participated in the poster session. Next MEM is scheduled to take place in two years in Mainz (making it a German Evolution Meeting or GEM). See you all there!
16.12.2024
Project Meeting "Ghost in the protein" in Prague

Last week we had our wonderful annual project meeting in Prague hosted by the group of Klara Hlouchova. Together with the groups of Florian Hollfelder from Cambridge and Ylva Ivarsson from Uppsala we work on a project funded by the Volkswagen foundation. On the photo are (from left to right) Klara, Margaux, Filip, Valerio, Lars, Priyanka, Erich, Isabella, Florian, Tea, Matus, Tereza, Matej, Bharat and Matthew. We are being watched over by a statue of Jan Huis, a famous czech theologian and former rector of Charles University.
09.12.2024
Bornberglab Christmas Party

We had our annual christmas party with an obligatory movie in the theater, this year How to Train your Dragon, a trip to the christmas market and a dinner with the whole group and students from the last year.
06.12.2024
Best PhD thesis prize for Lars

Very proud that our PhD student and current Postdoc, Lars Eicholt, received the Faculty’s Best PhD Thesis Award from the university’s rectorate. All the recipients and the summaries of their dissertations can be found here.
06.12.2024
Best MSc thesis prize in Evolutionary Biology for Marie

Delighted that our former MSc student and now PhD student, Marie Kristin Lebherz, received the annual "Bernhard-Rensch Prize for Evolution" for the best Master Thesis with a focus on Evolutionary Biology!
25.11.2024
EBB talk at EMBO in Greece

Erich gave a talk at the Evolutionary and Comparative Genomics lecture course in the beatuiful town of Nafplion, Greece. Thanks to Nikos for organising this course and inviting.
21.10.2024
Group retreat in Merano, South Tyrol

This year's group retreat was in beautiful Merano in Italy, where we were able to enjoy the mild and not at all rainy weather for a whole week. Our special guest again was Richard Goldstein from UCL, who gave not one, but two great talks this time! We also did a day trip to see the real Ötzi in Bolzano. More photos will follow here
05.09.2024
Bornberglab family reunion at OIST

Three of our lab members are currently visiting collaborators at OIST in Okinawa, Japan. Alina, Lars and Erich met in front of the Sydney Brenner statue who served as president for the institute from 2005 to 2011. Alina is working together with Tom Bourguignon while Lars and Erich are visiting Paola Laurino's lab.
03.09.2024
Talks on ICE (International Congress of Entomology)

Our PhD student Alina presented her research on the "Genomic basis of sociality: from cockroaches to eusocial termites" at the International Congress of Entomology (ICE) last week. ICE took place in Kyoto, Japan this year. Erich gave a talk about the research within the priority program "Genomic Basis of Evolutionary Innovations" (GEVol) funded by the DFG.
30.08.2024
New Paper! Sequence, Structure, and Functional Space of Drosophila De Novo Proteins
Lasse Middendorf, Bharat Ravi , Lars Eicholt Genome Biology and Evolution
In this new study, Lasse, Bharat and Lars explored the structural and functional diversity of de novo proteins by refining AlphaFold models for around 1,500 of them using molecular dynamics simulations. Most de novo proteins were found to be structurally flexible and did not resemble known folds, indicating they occupy a unique region in sequence space. However, a few showed high structural similarity to known protein domains, such as ancient folds like SH3 and HTH. They predicted Gene Ontology (GO) terms to assess their potential activities, revealing a wide range of functions, including transferase and hydrolase activity, and binding of nucleic acids and ions. The high degree of structural disorder and diverse GO terms led us to hypothesize that some de novo proteins might form biomolecular condensates. By clustering these proteins based on sequence features of intrinsically disordered regions, we identified 63 de novo proteins likely to form condensates, with older de novo proteins more enriched in this group. Interestingly, while most de novo proteins are closer to random sequences, a few align more closely with conserved proteins. This suggests that while most de novo proteins resemble random sequences, a subset has recognizable structures and functions. These findings provide a comprehensive view of de novo protein diversity and offer a foundation for more focused experimental studies in the future.
01.08.2024
Paper Alert! How antisense transcripts can evolve to encode novel proteins
Bharat Ravi , Anna Grandchamp , Erich Bornberg-Bauer Nature Communications
New protein coding genes can emerge de novo, overlapping existing protein genes but in the opposite (antisense) orientation. The authors investigated the possibility of such events using mathematical modelling and data analysis, and found that emergence of a protein coding region (ORF) is generally more likely in one frame of overlap (frame 1), than in the other two frames of overlap and in non-overlapping intergenic regions.
22.07.2024
Marie at SMBE in Mexico presenting her latest paper
Marie Lebherz, Bertrand Fouks, Julian Schmidt, Erich Bornberg-Bauer, Anna Grandchamp Genome Biology & Evolution
We are so proud of our PhD student Marie Lebherz who went to SMBE in Puerto Vallarta, Mexico and even gave a talk. Her presentation entitled "DNA Transposons favour de novo transcript emergence through enrichment of transcription factor binding motifs" was part of the symposium about "Transposable Elements in the population genomics era : towards a better understanding of their contribution to evolution". Transposable elements (TE) are known to play an essential role in driving evolvability and thus facilitating fast acting molecular adaptive processes at the DNA level. TEs can affect the recently detected and meanwhile well confirmed process of de novo gene emergence, specifically the gain of new transcription events. Overall, TE insertions play an important role in new transcript emergence, partly by introducing new regulatory motifs. For more details on the findings, we can now also happily announce that the paper has been recently published in GBE.
19.06.2024
New Positions for our postdocs

Exciting news for our group! Two of our postdocs will soon start their permanent positions in Universities across Europe. Anna Grandchamp (right in the picture) will soon be Maitre de conférence in Marseille. Mark Harrison (left in the picture) will be Associate Professor at Coventry University.
12.06.2024
PhD defence Margaux Aubel

Yesterday Margaux successfully defended her PhD on the "Evolution of structural properties and folding in human de novo proteins". Until the end of this year she will stay in the group and continue working on exciting projects about protein evolution.
03.06.2024
Paper out: Complete Sex Chromosomes of Great Apes sequenced... And there are de novo genes!
Kateryna D. Makova, Brandon D. Pickett, Robert S. Harris, [...], Margaux Aubel, Erich Bornberg-Bauer, [...], Adam M. Phillippy Nature
Last week the complete X and Y chromosomes from living great apes were published lead by the Makova Lab. We are very happy to have contributed a small analysis on de novo genes in this huge and important study. We were able to detect de novo genes on the highly repetitive and fast evolving Y chromosomes of some of the great apes. One of the de novo genes (shown in the figure) had been duplicated many times in bonobo, similar to the highly duplicated Y chromosome-specific genes called ampliconic genes. For a nice summary of the complete study including statements from the leading authors take a look at the original press release from the Penn State University.
31.05.2024
Paper out in Journal of Evolutionary Biology! - Major changes in domain arrangements are associated with the evolution of termites
Alina Mikhailova, Elias Dohmen, Mark Harrison Journal of Evolutionary Biology
Proteins have functional units called domains, and their rearrangements can reveal changes linked to complex traits like eusociality. In termites, which evolved eusociality from cockroaches, there was a reduction in proteome size, suggesting they repurposed existing proteins. We studied domain rearrangements in three cockroaches and five termites and found over 5,000 such changes. At the origin of termites, 90 new domain arrangements were linked to longevity-related functions, crucial for new castes. As termite societies became more complex, with sterile workers and larger colonies, 110 more domain arrangements emerged, related to protein glycosylation and ion transport. These genes often showed caste-specific expression and higher DNA methylation. Our findings highlight the importance of domain rearrangements in evolving termite eusociality.
17.05.2024
PhD defence Lars Eicholt

This week Lars successfully defended his PhD. In his thesis he looked at "Structural Analysis of de novo emerged proteins". Luckily he will stay in the group until the end of the year and continue his enterprises on the structural characterisation of de novo proteins.
22.04.2024
Official Graduation of Brennen and Amer

At the graduation last Friday Brennen Heames and Amer Ghalawinji officially received their doctoral degrees. Brennen, who was part of our own group, came all the way back from the UK to receive his degree. He investigated the birth of de novo genes in Drosophila and looked more closely at their structural properties in comparison to random proteins. Amer from Prof. Monika Stoll's group looked at as part of the EvoPad graduate schoolthe effect of lncRNAs on genetic predisposition to heart failure with Erich as co-supervisor. Congratulations to both of them!
18.04.2024
Paper out in Genome Biology and Evolution!
Margaux Aubel, Filip Buchel, Brennen Heames, Alun Jones, Ondrej Honc, Erich Bornberg-Bauer, Klara Hlouchova Genome Biology and Evolution
Together with our collaborators Klara Hlouchova, Filip Buchel and Ondrej Honc in Prague, we looked into the folding propensities of de novo emerged proteins. Akin to random sequences, proteins that emerge de novo from noncoding DNA are usually predicted to be highly disordered as they inherit no structure from their ancestor, unlike proteins evolving through duplication. Most studies to-date have relied on computational prediction of the structural properties of de novo proteins. Here, we show experimentally that, while most of our putative human de novo proteins are highly disordered, some contain propensity for structure and globularity, seen most clearly in the oldest de novo proteins in our dataset. With that we aim to lay the groundwork for experimental verification of hypotheses regarding the structural evolution of de novo proteins.
20.01.2024
Paper out in Proteins!
Lasse Middendorf, Lars Eicholt Proteins: Structure, Function and Bioinformatics
The paper, "Random de novo and Conserved Proteins: How Structure and Disorder Predictors Perform Differently," by Lasse and Lars, investigates the structural characteristics of de novo, random, and conserved proteins using protein structure prediction tools like AlphaFold2 and ESMFold. It examines how these tools perform with such proteins lacking sequence identity and highlights the differences in structural predictions and confidence scores (pLDDT) between these protein. The study also explores the influence of sequence length and Multi-Sequence Alignment (MSA) depth on prediction outcomes. The findings suggest a unique structural and disorder profile for de novo and random proteins in which pLDDT correlates negatively with beta-sheets and positively with disorder and alpha-helices. In conserved proteins pLDDT normally correlates negatively with disorder and is often used as an indicator of disordered regions.
12.01.2024
Master Thesis Defense Of Alina and Lasse

Starting into the New Year with not only one, but two great Master Thesis defenses! First Alina Berger defended her thesis on the "Evolution of the CatSpermasome". She looked into the evolutionary history of the sperm-specific ion channel, which is essential for male fertility to track down its emergence throughout vertebrates and detected signatures of positive selection along the phylogenetic tree. Her second thesis advisor was Timo Strünker from the UKM.
Then Lasse Middendorf presented and successfully defeded his thesis on "
4.12.2023
PhD defence Brennen Heames

Last friday we had the honour to listen to Brennen's PhD defence about "Exploring the potential of de novo gene emergence to support protein-coding evolutionary novelty". It was a great talk and it has been a pleasure to have him in the group!
10.11.2023
Anna and Erich in Texas

Erich and Anna represented the group at the SMBE Sattelite Meeting on "De novo Gene Birth" in College Station, Texas. The meeting was beautifully organised by Claudio Casola, Li Zhao, Victor Luria and Nikolaos Vakirlis. We are looking forward to seeing you all again, hopefully at the next SMBE in Puerto Vallarta, Mexico next July.
24.10.2023
Biospektrum Article on de novo Proteins

The experimentalists of our group Margaux, Lars and Andreas wrote an article with Erich for the german magazine Biospektrum about our work on de novo proteins. In the article the focus is mainly on the experimental side of the research, which is explained to a broad audience not familar to de novo gene evolution.
30.07.2023
SMBE in Italy

The SMBE 2023 took place in the beautiful Ferrara in Italy. Erich and Claudia Alvarez-Carreño organised the symposium on "Novel proteins and their emergence from LUCA until today" where Lars gave his talk. Elias presented his work in the Symposium on "Computational evolutionary genomics in the era of machine learning" and Margaux as part of the "Graduate Students Excellence Awards" Symposium. Bharat presented his poster in "Evolutionary biology through a functional genomics lens". We hope to see many people again next year in Mexico or at the SMBE Sattelite Meeting on "De novo Gene Birth" in Texas in November!
25.07.2023
Population genomics reveals mechanisms and dynamics of de novo expressed open reading frame emergence in Drosophila melanogaster
Anna Grandchamp, Lucas Kühl, Marie Lebherz, Kathrin Brüggemann, John Parsch, Erich Bornberg-Bauer Genome Research

Congratulations Anna and Marie on this really cool new paper! They found that novel genes are essential for evolutionary innovations and that they differ substantially even between closely related species. Some of those novel genes arise de novo, that is, from previously noncoding DNA. To characterize the underlying mutations that allowed de novo gene emergence and their order of occurrence, homologous regions must be detected within noncoding sequences in closely related sister genomes. Here they sequenced and assembled genomes with long-read technology and the corresponding transcriptomes from inbred lines of Drosophila melanogaster, derived from seven geographically diverse populations (see picture on the left). The new data suggests a rapid turnover of novel transcripts and more frequent gain and loss of transcription than the creation of ORFs. The highly mutable genomic regions around TEs provide new features that enable gene birth. In conclusion, de novo ORFs have a high birth-death rate, are rapidly purged, but surviving de novo ORFs spread neutrally through populations and within genomes.
13.07.2023
Protein Evolution Meeting Münster

We had a great Protein Evolution Meeting in Münster (PEMM) this week! Thanks to all speakers, visitors, poster creators and helpers. We hope to see everyone again soon and look forward to the next meeting of this kind.
10.06.2023
University News

Great article about our group and our research on de novo proteins from our university newspaper! Check it out here.
18.04.2023
Neutral models of de novo gene emergence suggest that gene evolution has a preferred trajectory
Bharat Ravi, Erich Bornberg-Bauer Molecular Biology and Evolution

Cheers and congratulations to Bharat! His new paper is about the theory behind the emergence of de novo genes.
New protein coding genes can emerge from genomic regions that previously did not contain any genes, via a process called de novo gene emergence. To synthesize a protein, DNA must be transcribed as well as translated. Both processes need certain DNA sequence features. Stable transcription requires promoters and a polyadenylation signal, while translation requires at least an open reading frame (ORF). We develop mathematical models based on mutation probabilities, and the assumption of neutral evolution, to find out how quickly genes emerge and are lost. We also investigate the effect of the order by which DNA features evolve, and if sequence composition is biased by mutation rate. We rationalize how genes are lost much more rapidly than they emerge, and how they preferentially arise in regions that are already transcribed. Our study not only answers some fundamental questions on the topic of de novo emergence but also provides a modeling framework for future studies.
07.04.2023
Experimental characterization of de novo proteins and their unevolved random-sequence counterparts
Brennen Heames, Filip Buchel, Margaux Aubel, Vyacheslav Tretyachenko, Dmitry Loginov, Petr Novák, Andreas Lange, Erich Bornberg-Bauer, Klara Hlouchova Nature Ecology & Evolution

Together with the KHlab in Prague we published our study on de novo proteins compared to random proteins. De novo gene emergence provides a route for new proteins to be formed from previously non-coding DNA. Proteins born in this way are considered random sequences and typically assumed to lack defined structure. Taking putative de novo proteins identified in human and fly, we experimentally characterize a library of these sequences to assess their solubility and structure propensity. We compare this library to a set of synthetic random proteins with no evolutionary history. Bioinformatic prediction suggests that de novo proteins may have remarkably similar distributions of biophysical properties to unevolved random sequences of a given length and amino acid composition. However, upon expression in vitro, de novo proteins exhibit moderately higher solubility which is further induced by the DnaK chaperone system. We suggest that while synthetic random sequences are a useful proxy for de novo proteins in terms of structure propensity, de novo proteins may be better integrated in the cellular system than random expectation, given their higher solubility.
06.04.2023
HFSP Awards 2023

Happy to announce that the lab just received its 4th HFSP grant! “New Kids on the Block: how De Novo emerged micropeptides rewire cellular networks”. This time with Anne-Ruxandra Carvunis and Christine Brun. HFSP Awards 2023
21.03.2023
Münster Evolution Meeting

Last week our group participated in the second Münster Evolution Meeting (MEM). There were many interesting talks and great posters and we were lucky to meet so many great people. It was a nice conference and many of us had the opportunity to present their research in a talk (like Anna Grandchamp here in the picture) or during the poster sessions. We want to thank the organising team from the Gadau group again and hope to see everyone again at the next MEM!
28.02.2023
Guest Researcher Aleksandra Marsavelski

For one exciting month we have a guest researcher from Zagreb, Croatia in our group. Aleksandra Marsavelski is visiting us as a WiRe fellow to look at plastic degrading enzymes. Together with our group she wants to reconstruct ancestors of enzymes with a known plastic degrading activity and enhance their function. This can help us in the future to recycle plastics in an environmentally friendly way. For more information and news, check out the WiRe website of the University here.
03.02.2023
Introducing creative destruction as a mechanism in protein evolution
Margaux Aubel, Erich Bornberg-Bauer Proceedings of the National Academy of Sciences

In this commentary we discuss the recent paper from Alvarez-Carreño et al. about the evolution of new folds. In particular, we look at ancient folds from the dawn of the RNA-protein world and how new and still existing folds may have originated through reshaping of the old folds. The mechanism introduced by Alvarez-Carreño et al., called "creative Destruction" describes the process of innovation through the “destruction” of an old product and thereby generating a new one. An analogy from everyday life is the development of an mp3 player from a cassette player. The old cassette player, lost its overall form and the need of a cassette tape to develop into an mp3 player. Form and function of both products are similar while some aspects of the ancestral product have been ‘destroyed’, i.e. cassette, bulky structure and loudspeaker, to create the new features, i.e. memory space, small, headphones.
01.02.2023
Blattodea Conference in April

27.01.2023
Kai's Master Thesis handed in and defended

Our Master student Kai has succesfully defended his Thesis last December on "Structural analysis of putative de novo protein Atlas and its ancestors". After mastering many obstacles in the wet lab, Kai found his true calling during the computational analyses. Now he is heading off to new computational adventures and we wish him all the best of luck on his journey!
07.12.2022
IEB seminar by Birte Höcker

Birte Höcker from the University of Bayreuth gave an IEB seminar on the Evolution and Design of Protein Folds. Thanks for visiting our group and the IEB!
01.12.2022
Life? Symposium in Hanover - Ghost in the protein

We had a great symposium at the Xplanatorium in Hanover Herrenhausen from the 29th to 30th of November. Together with our collaborators Ylva Ivarsson from Uppsala, Florian Hollfelder from Cambridge and Klara Hlouchova from Prague we look into the early evolution of proteins from random sequence space. Our project on the emergence of new proteins by de novo mechanism is funded by the Volkswagen Stiftung Life? initiative together with several other interesting projects.
30.11.2022
2nd Münster Evolution Meeting from 13th-16th March 2023. Registration is open!

We are happy to announce that MEM is finally back! See you all in Münster from 13th -16th March 2023. The Münster Evolution Meeting (MEM) aims to provide a forum for all Evolutionary Biologists working across different fields. Besides having the opportunity to share and learn about excellent research in evolutionary biology MEM also aims at bringing together Evolutionary Biologists working in German-speaking countries in a smaller setting, to allow for intensive networking and discussion. Münster is a welcoming and vibrant university town, offering a perfect venue. It was the home of Prof. Dr. Bernhard Rensch, who contributed significantly to the “Modern Evolutionary Synthesis”. Click here to register!
03.11.2022
Eusocial Transition in Blattodea: Transposable Elements and Shifts of Gene Expression
Juliette Berger, Frédéric Legendre, Kevin-Markus Zelosko , Mark C. Harrison, Philippe Grandcolas, Erich Bornberg-Bauer, Bertrand Fouks Genes

Using available genome and transcriptome of cockroaches and termites as well as de novo transposon annotations, our study brings further evidence on the role of transposon-aided adaptations during termite eusocial transition. Our study reveals that transposon insertions within differentially expressed genes (DEGs) among castes in termites and life-stages (corresponding to termite castes) in cockroaches display opposite patterns. Furthermore, while in cockroaches transposon families do not differ in their insertion frequency within DEGs, retrotransposon and DNA transposon insertions are more frequent in worker- and queen-biased genes, respectively. In addition, the insertion of transposons in queen-biased genes in termites are more frequent in genes involved in reproduction, ageing, and immunity, functions essential for queen phenotype; while transposon insertion in termite worker-biased genes are more frequent in genes involved in the regulation of gene expression and behaviour, functions associated with phenotypic plasticity and the apparition of a sterile caste. In conclusion, our study highlights the role of transposon during the eusocial transition in Blattodea, potentially through shifts in gene expression.
12.10.2022
Sarah's Master Thesis Celebration & PhD start

After Sarah's successful Master thesis defence we had some delicious Flammkuchen together with her family. The title of her thesis was "The role of selection in the emergence of caste-biased genes in termites" and she was under the amazing supervision of Mark. We are very happy that she now started her PhD in the group on the evolutionary genomics of sociality in beetles funded by GEvol (SPP 2349)
11.10.2022
Domain Evolution of Vertebrate Blood Coagulation Cascade Proteins
Abdulbaki Coban, Erich Bornberg-Bauer, Carsten Kemena Jornal of Molecular Evolution

Here we examined proteomes of 21 chordates, of which 18 are vertebrates, to reveal the modular evolution of the blood coagulation cascade. Within the vertebrate coagulation protein set, almost half of the studied proteins are shared with jawless vertebrates. Domain similarity analyses revealed that there are multiple possible evolutionary trajectories for each coagulation protein. During the evolution of higher vertebrate clades, gene and genome duplications led to the formation of other coagulation cascade proteins.
04.10.2022
Group retreat in Pfaffenhofen a. d. Roth

We just got back from our awesome second group retreat this year! A lot of great presentations and talks, especially from and with our special guest Richard Goldstein. Apart from the serious scientific topics, we went for a beautiful walk to see the famous "Küssende Sau" (kissing pig). Stay tuned for more pictures...
29.09.2022
Priority Programme “Genomic Basis of Evolutionary Innovations (GEvol)” (SPP 2349) kick-off meeting in Münster
Official website of the SPP GEvol

The kick-off meeting for the DFG Priority Programme “Genomic Basis of Evolutionary Innovations (GEvol)” (SPP 2349) coordinated by Erich was the first in-person meeting of nearly all of the 40 involved PIs! The goal of the Priority Programme is to exploit new methods to reveal in the insect taxon the role of: coding vs. regulatory changes, transposable elements, epigenetic regulation, gene family evolution, copy number dynamics, structural genomic rearrangements etc. in trait evolution by using multiple cutting edge quantitative OMICs resources (e.g. genomics, transcriptomics and epigenomics). Eventually, the emerging hypotheses shall be tested by functional genetics experiments where possible.
13.09.2022
Vector redesign and in-droplet cell-growth improves enrichment and recovery in live Escherichia coli
Bernard D. G. Eenink, Tomasz S. Kaminski, Erich Bornberg-Bauer, Joachim Jose, Florian Hollfelder, Bert van Loo Microbial Biotechnology

In this article we compare different methods to sort large libraries of enzymes presented on the surface of E. coli, and recovering the intact cells without lysis. We changed a protocol in which single cells encapsulated in a microdroplet are measured using Fluorescence-activated droplet sorting (FADS) to a protocol were cells are grown inside a microdroplet after encapsulation of a single cell, resulting in multiple copies of the same variant inside each droplet. We find that growing cells inside a microdroplet prior to FADS increases both the amount of cells recovered as well as reducing the amount of false positives.
30.08.2022
The role of transposable elemets in the evolution of insect socal complexity - Erich Bornberg-Bauer at the HFSP meeting in Paris

This time the group leader Erich Bornberg-Bauer himself presented our work on the role of transposable elements in termites. The 21st HFSP Awardees meeting took place in beautiful Paris this year. The poster is based on work by postdoc Mark Harrison and former postdoc Evelien Jongepier from our group in cooperation with Mireille Vasseur-Cognet (Paris), Hei Sook Sul (Berkeley) and Wilhelm de Beer (Pretoria).
We compared whole-genome levels of repeat element transcription in the fat body of female workers, kings, and five different
queen stages. Those queens can live over 20 years, maintaining near maximum reproductive output! The sterile workers
only live weeks or months. We found substantially higher TE activity in workers than in
reproductives. Furthermore, relative TE expression decreased with age in queens, due
to a significant upregulation of the PIWI-pathway in 20-year-old queens. Our results
suggest a caste- and age-specific regulation of the PIWI-pathway has evolved in higher
termites that is analogous to germ-line-specific activity in individual organisms,
promoting reproductive fitness even at high age.
30.08.2022
Evidence for a conserved queen-worker genetic toolkit across slave-making ants and their ant hosts
Barbara Feldmeyer, Claudia Gstöttl, Jennifer Wallner, Evelien Jongepier, Alice Séguret, Donato A. Grasso, Erich Bornberg-Bauer, Susanne Foitzik, Jurgen Heinze Molecular Ecology

The ecological success of social Hymenoptera (ants, bees, wasps) depends on the division of labour between the queen and workers. Each caste exhibits highly specialized morphology, behaviour, and life-history traits, such as lifespan and fecundity. Despite strong defences against alien intruders, insect societies are vulnerable to social parasites, such as workerless inquilines or slave-making ants. Here, we investigate whether gene expression varies in parallel ways between lifestyles (slave-making versus host ants) across five independent origins of ant slavery in the “Formicoxenus-group” of the ant tribe Crematogastrini. As caste differences are often less pronounced in slave-making ants than in nonparasitic ants, we also compare caste-specific gene expression patterns between lifestyles. We demonstrate a substantial overlap in expression differences between queens and workers across taxa, irrespective of lifestyle. Caste affects the transcriptomes much more profoundly than lifestyle, as indicated by 37 times more genes being linked to caste than to lifestyle and by multiple caste-associated modules of coexpressed genes with strong connectivity. However, several genes and one gene module are linked to slave-making across the independent origins of this parasitic lifestyle, pointing to some evolutionary convergence. Finally, we do not find evidence for an interaction between caste and lifestyle, indicating that caste differences in gene expression remain consistent even when species switch to a parasitic lifestyle. Our findings strongly support the existence of a core set of genes whose expression is linked to the queen and worker caste in this ant taxon, as proposed by the “genetic toolkit” hypothesis.
23.08.2022
Anna Grandchamp and Alina Mikhailova present their work at ESEB in Prague

Our postdoc Anna Grandchamp presented her poster on proto-gene emergence and PhD student Alina Mikhailova gave a talk about domain rearrangements in social termites. Awesome work by both of them!
21.07.2022
MC Harrison on ICE (International Congress of Entomology)

Our postdoc Mark C Harrison presented his work on social insects this week on the International Congress of Entomology (ICE) in Helsinki, Finnland. Check out his latest publications in Open Biology and Genome Biology and Evolution
21.07.2022
Complex regulatory role of DNA methylation in caste- and age-specific expression of a termite
Mark C. Harrison, Elias Dohmen, Simon George, David Sillam-Dussès, Sarah Séité, Mireille Vasseur-Cognet Open Biology

The reproductive castes of eusocial insects are often characterized by extreme lifespans and reproductive output, indicating an absence of the fecundity/ longevity trade-off. The role of DNA methylation in the regulation of caste- and age-specific gene expression in eusocial insects is controversial. While some studies find a clear link to caste formation in honeybees and ants, others find no correlation when replication is increased across independent colonies. Although recent studies have identified transcription patterns involved in the maintenance of high reproduction throughout the long lives of queens, the role of DNA methylation in the regulation of these genes is unknown. We carried out a comparative analysis of DNA methylation in the regulation of caste-specific transcription and its importance for the regulation of fertility and longevity in queens of the higher termite Macrotermes natalensis. We found evidence for significant, well-regulated changes in DNA methyl- ation in mature compared to young queens, especially in several genes related to ageing and fecundity in mature queens. We also found a strong link between methylation and caste-specific alternative splicing. This study reveals a complex regulatory role of fat body DNA methylation both in the div- ision of labour in termites, and during the reproductive maturation of queens.
13.07.2022
Heterologous expression of naturally evolved putative de novo proteins with chaperones
Lars A. Eicholt, Margaux Aubel, Katrin Berk, Erich Bornberg-Bauer, Andreas Lange Protein Science

Today, we know that proteins do not only evolve by duplication and divergence of existing proteins but also arise from previously non-coding DNA. These proteins are called de novo proteins. Their properties are still poorly understood and their experimental analysis faces major obstacles. Here, we aim to present a starting point for soluble expression of de novo proteins with the help of chaperones and thereby enable further characterization.
Over the past decade, evidence has accumulated that new protein-coding genes can emerge de novo from previously non-coding DNA. Most studies have focused on large scale computational predictions of de novo protein-coding genes across a wide range of organisms. In contrast, experimental data concerning the folding and function of de novo proteins are scarce. This might be due to difficulties in handling de novo proteins in vitro, as most are short and predicted to be disordered. Here, we propose a guideline for the effective expression of eukaryotic de novo proteins in Escherichia coli. We used 11 sequences from Drosophila melanogaster and 10 from Homo sapiens for heterologous expression. The candidate de novo proteins have varying secondary structure and disorder content. Using multiple combinations of purification tags, E. coli expression strains, and chaperone systems, we were able to increase the number of solubly expressed putative de novo proteins from 30% to 62%. We found that, overall, proteins with higher predicted disorder were easier to express.
26.04.2022
Janina (GadauLab) and Lars elected as IEB student representatives

Our PhD student Lars and Janina Rinke from the lab of Jürgen Gadau were elected as IEB student representatives for the next two years!
26.04.2022
Master Defence Christopher Finke: "Genome annotation and comparison of genomic features between slave-maker ants and their hosts"

Our dear student Christopher Finke has successfully defended his Master thesis yesterday evening. Supervised by Alice Séguret he looked into differences between slave-maker ants and their host species and was able to generate valuable genome annotations. Christopher started out as a Bachelor student in the lab with us and moved on to computational work during his Master studies. Congratulations and good luck in Berlin!
21.04.2022
Plant biodiversity assessment through soil eDNA reflects temporal and local diversity
María Ariza, Bertrand Fouks, Quentin Mauvisseau, Rune Halvorsen, Inger Greve Alsos, Hugo de Boer Methods in Ecology and Evolution

The use of environmental DNA as a proxy to identify species has increased exponentially in the last few years. However, how good eDNA relates with actual species presence/abundance remains elusive. Our study fill this gap, comparing both soil eDNA and visual assessments of plants in Norway. It shows that soil eDNA allows to describe well local plant biodiversity, with an average 60% match with visual survey for vascular plants, and also to recover past vegetation biodiversity. Our soil eDNA method then provide a great tool to assess rapidly plant biodiversity in any seasons.
08.04.2022
PhD students Margaux and Lars presenting their posters at Apfed22 in Bayreuth

Our PhD students Margaux Aubel and Lars Eicholt presented their most recent work on soluble expression of putative de novo proteins at Apfed22 in Bayreuth! If you did not attend, please check out their preprints here and here. We would like to thank all the organizers for this amazing hybrid conference. The speaker line-up was diverse both in backgrounds and research areas, while reaching gender parity!
22.02.2022
Co-expression analyses, combined with lipidomics and metabolomics, uncover lifespan prolonging mechanisms in extremely long-lived and highly fertile termite queens
Sarah Séité, Mark C Harrison, David Sillam-Dussès, Roland Lupoli, Tom J M Van Dooren, Alain Robert, Laure-Anne Poissonnier, Arnaud Lemainque, David Renault, Sébastien Acket, Muriel Andrieu, José Viscarra, Hei Sook Sul, Z Wilhelm de Beer, Erich Bornberg-Bauer , Mireille Vasseur-Cognet Communications Biology
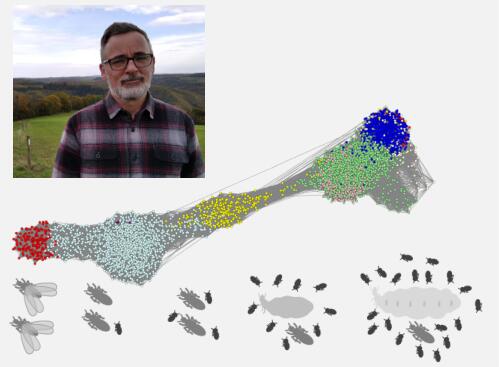
Some insects are eusocial, which means that their society is organized into different castes which carry out specific colony tasks. In termites, for example, the “king” and the “queen” are involved in reproduction, while the “workers” are involved in resource gathering and brood-care. Macrotermes natalensis is a species of higher termite that creates large complex colonies, in which the king and the queen (reproductives) have a long life, spanning decades, and the queen remains highly fertile throughout her adult life. Workers, on the other hand, are short-lived and sterile. We studied this species of termites, using a combination of high throughput techniques such as transcriptomics, metabolomics and lipidomics, to identify mechanisms that allow reproductives to live orders of magnitude longer than workers, while maintaining high fertility. Aging associated genes were differentially expressed between the reproductives and the workers. For example, antioxidant genes were highly expressed, and the membrane lipids were less damaged by oxidative stress, in the queens relative to workers. Contrary to expectations, we found that several members of the insulin/insulin-like growth factor signaling (IIS) pathway were upregulated in the queens. Normally this would indicate increased diversion of metabolism towards energy storage. However, we did not find excessive fat storage in the queens; simple sugars dominate in their hemolymph and a large amount of resources is allocated to egg production. Our findings support the notion that aging results from a complex interplay of several processes. These processes should be studied simultaneously and not in isolation, for a better understanding of aging.
20.10.2021
Convergent loss of chemoreceptors across independent origins of slave-making in ants

Socially parasitic ants exploit the work force and social organisation of closely related ant species, and therefore no longer need to perform certain social tasks such as brood care and foraging. The loss of such behaviours is expected to be accompanied by loss of the underlying genes. We sequenced the genomes of eight ant species representing three independent origins of parasitism. Due to their importance in chemical communication and foraging, we investigated the evolution of chemoreceptors in parasites and their hosts. We found that parasites lost a striking 50% of their gustatory receptor repertoire compared to their hosts, perhaps reflecting the outsourcing of foraging tasks to host workers. We also found that parasites had fewer olfactory receptors than their hosts, with the same olfactory receptors being lost across multiple origins of parasitism. This represents a rare case of convergent molecular evolution at the level of individual genes, shedding light on the loss of important social traits during the transition to a parasitic lifestyle.
12.10.2021
Ancestral sequences of a large promiscuous enzyme family correspond to bridges in sequence space in a network representation

Evolutionary relationships of protein families can be characterized either by networks or by trees. Whereas trees allow for hierarchical grouping and reconstruction of the most likely ancestral sequences, networks lack a time axis but allow for thresholds of pairwise sequence identity to be chosen and, therefore, the clustering of family members with presumably more similar functions. Here, we use the large family of arylsulfatases and phosphonate monoester hydrolases to investigate similarities, strengths and weaknesses in tree and network representations. For varying thresholds of pairwise sequence identity, values of betweenness centrality and clustering coefficients were derived for nodes of the reconstructed ancestors to measure the propensity to act as a bridge in a network. Based on these properties, ancestral protein sequences emerge as bridges in protein sequence networks. Interestingly, many ancestral protein sequences appear close to extant sequences. Therefore, reconstructed ancestor sequences might also be interpreted as yet-to-be-identified homologues. The concept of ancestor reconstruction is compared to consensus sequences, too. It was found that hub sequences in a network, e.g. reconstructed ancestral sequences that are connected to many neighbouring sequences, share closer similarity with derived consensus sequences. Therefore, some reconstructed ancestor sequences can also be interpreted as consensus sequences.
30.03.2021
New research priorities in Münster

Die Deutsche Forschungsgemeinschaft (DFG) fördert zwei neue Schwerpunktprogramme (SSP) an der Westfälischen Wilhelms-Universität in Münster. Insgesamt fließen dafür in den nächsten drei Jahren 10 bis 14 Millionen Euro nach Münster. Gefördert werden Programme in der Biologie und der Chemie. Im Projekt „Die genomischen Grundlagen evolutionärer Innovationen (GEvol)“ unter der Leitung des Biologen Prof. Dr. Erich Bornberg-Bauer vom Institut für Evolution und Biodiversität wollen die Forscher die Mechanismen der genetischen Veränderungen einer großen Artengruppe entschlüsseln. „In dem Vorhaben untersuchen wir unter anderem die Prozesse, die den wichtigsten genomischen Veränderungen in der Evolution zugrunde liegen – beispielsweise Gewinn und Verlust von Sozialität oder Paarungssystemen, Verteidigung und Immunität, entwicklungsbiologischen und morphologischen Anpassungen“, sagt Erich Bornberg-Bauer. Read more
29.03.2021
Münster University receives two new research associations

The University of Münster is coordinating two new Priority Programmes funded by the German Research Foundation (DFG) with several million euros. The projects come from the fields of biology and chemistry and deal with innovative informatics technologies.Read more
12.03.2021
Structural and functional characterization of a putative de novo gene in Drosophila
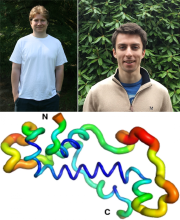

Comparative genomic studies have repeatedly shown that new protein-coding genes can emerge de novo from noncoding DNA. Still unknown is how and when the structures of encoded de novo proteins emerge and evolve. Combining biochemical, genetic and evolutionary analyses, we elucidate the function and structure of goddard, a gene which appears to have evolved de novo at least 50 million years ago within the Drosophila genus. Previous studies found that goddard is required for male fertility. Here, we show that Goddard protein localizes to elongating sperm axonemes and that in its absence, elongated spermatids fail to undergo individualization. Combining modelling, NMR and circular dichroism (CD) data, we show that Goddard protein contains a large central α-helix, but is otherwise partially disordered. We find similar results for Goddard’s orthologs from divergent fly species and their reconstructed ancestral sequences. Accordingly, Goddard’s structure appears to have been maintained with only minor changes over millions of years.
12.03.2021
New proteins 'out of nothing'

Proteins are the key component in all modern forms of life. Haemoglobin, for example, transports the oxygen in our blood; photosynthesis proteins in the leaves of plants convert sunlight into energy; and fungal enzymes help us to brew beer and bake bread. Researchers have long been examining the question of how proteins mutate or come into existence in the course of millennia. That completely new proteins - and, with them, new properties - can emerge practically out of nothing, was inconceivable for decades, in line with what the Greek philosopher Parmenides said: "Nothing can emerge from nothing" (ex nihilo nihil fit). Working with colleagues from the USA and Australia, researchers from the University of Münster (Germany) have now reconstructed how evolution forms the structure and function of a newly emerged protein in flies. This protein is essential for male fertility. The results have been published in the journal "Nature Communications". Read more
19.02.2021
Marie Skłodowska-Curie Actions | Individual Fellowship awarded to Dr. Bertrand Fouks

How genomes evolve and drive novelty is a central question in biology. Some of the most puzzling genomic innovations, for example the development of placenta in mammals, are triggered by Transposable Elements (TEs). TEs are small genome fragments that can move and insert in other areas of the genome, which can create or impair gene functions. Organisms have adapted mechanisms to counteract the harmful effects of TEs, notably small RNAs (e.g. piwi-interacting RNA, piRNAs). Despite increasing knowledge on the effects of TEs on genome evolution and the apparition of novel traits, how and which TEs along with their interactions with piRNAs can promote novelty remain unclear. The project of Dr. Fouks will shed light on this issue by investigating how TEs and piRNAs evolved and interacted in cockroaches and termites alongside the evolution of their incredible biodiversity, with an emphasis on sociality and wood feeding. Dr. Fouks will generate several high-resolution genomes and transcriptomes from cockroach and termite species to locate and categorize TEs and piRNAs., This will allow him to unravel their role in the adaptation of cockroaches and termites to different social levels and diets.
24.07.2020
Humboldt Fellowship for Dr. Anna Grandchamp

Since several years it is known that new proteins not only arise via gene duplication and variation of the duplicates but also de novo, i.e. from previously non-coding DNA. An important first step in the creation of these de novo genes is that some of the zillions of randomly generated transcripts have some, though very weak, inherent function or are at least not toxic to the cell and are not quickly lost again. In her project, Dr. Grandchamp will investigate how often new random transcripts are created, by which mechanisms they are created and what the initial function of the new proteins might be.She plans to use in-bred lines of fly populations collected from all over Europe as well as of closely related fly species and map their transcriptomes onto the newly sequenced genomes to precisely characterise the creation and loss of de novo genes.
22.10.2019
Researchers gain new insights into the evolution of proteins

How do bacteria manage to adapt to synthetic environmental toxins and to even develop strategies for using a pesticide agent as food within less than 70 years? This is what scientists at Münster University have investigated. They found out how mutations led to biochemical changes that now enable an enzyme to cleave a pesticide. The study was published in "Nature Chemical Biology". Read more
11.09.2018
Bioinformaticians examine new genes the moment they are born

Accumulating evidence suggests that new genes can arise spontaneously from previously non-coding DNA instead of through the gradual mutation of established genes. Bioinformaticians at the University of Münster are now, for the first time, studying the earliest stages in the emergence of such “genes out of thin air”, also known as de novo genes. Read more
26.04.2018
"The Elusive Calculus of Insect Altruism

In 1964, the evolutionary biologist William D. Hamilton seemingly explained one of the greatest paradoxes in biology with a simple mathematical equation. Even Charles Darwin had called the problem his “one special difficulty” a century earlier in On the Origin of Species, writing that it made him doubt his own theory. The paradox in question is the altruistic behavior exhibited most famously by social insects. Ants, termites, and some bees and wasps live in highly organized colonies in which most individuals are sterile or forgo reproduction, instead serving the select few who do lay eggs. Yet such behavior seemed to clearly violate the concept of natural selection and survival of the fittest, if “fittest” means the individual with the greatest reproductive success. The insects’ compulsory altruism—a form of extreme social behavior called eusociality—made little sense. Read more
06.04.2018
"Human Frontier Science Program" funds two projects involving Münster University researchers

In the selection phase for 2018, the prestigious “Program Grant” research award given by the international “Human Frontier Science Program” goes to two members of the Department of Biology at the University of Münster – to bioinformatics specialist Prof. Erich Bornberg-Bauer and cell biologist Prof. Karin Busch. Read more
22.03.2018
Consider the Cockroach

On the fringes of urban life, some creatures thrive more than others. The brown rat, the coyote, the Canada goose, the Northern raccoon and many other species have all expanded their ranges and populations because they are well-adapted to scavenge or hunt in the shadows of human civilization. But perhaps none have been as successful as the cockroach. Read more
07.02.2018
The social evolution of termites
One phenomenon that already fascinated Charles Darwin is the evolution of huge, complex insect societies from solitary ancestors. This was the case with termites and ants, which have the same eusocial lifestyle. This has distinctive features such as the creation of castes, including for instance a complex system of division of labor among workers and soldiers. A team headed by evolutionary biologist Prof. Dr. Judith Korb from the University of Freiburg, bioinformatician Prof. Dr. Erich Bornberg-Bauer, evolutionary biologist Dr. Mark Harrison and evolutionary biologist Dr. Evelien Jongepier from the Westphalian Wilhelms University in Münster has now compared the molecular basis for the evolution of the eusocial lifestyle. Read more
05.02.2018
Scientists investigate the molecular basis of social evolution in Termites

Researchers from the group of bioinformatician Prof Erich Bornberg-Bauer from the Institute for Evolution and Biodiversity at the WWU have now, for the first time, compared the molecular basis for the evolution of eusociality within termites and ants. Read more
01.02.2016
Land plant became key marine species
The genome of eelgrass (Zostera marina) has now been unveiled. It turns out that the plant, once land-living but now only found in the marine environment, has lost the genes required to survive out of the water. Scientists from the University of Gothenburg participated in the research study, the results of which are published in the scientific journal Nature. Eelgrass belongs to a group of flowering plants that have adapted to a life in water. As such, it is a suitable candidate for studies of adaptation and evolution. 'Since flowering plants have emerged and developed on land, eelgrass can be expected to share many genetic features with many land plants. Studying differences between them can tell us how eelgrass has adapted to a marine environment,' says Mats Töpel, researcher at the Department of Marine Sciences, University of Gothenburg, who participated in the sequencing of the eelgrass genome. Töpel is part of an international research collaboration involving 35 research teams. As a result of their efforts, the eelgrass genome has now been published in Nature. Read more
31.08.2015
A Surprise Source of Life's Code

Genes, like people, have families — lineages that stretch back through time, all the way to a founding member. That ancestor multiplied and spread, morphing a bit with each new iteration. For most of the last 40 years, scientists thought that this was the primary way new genes were born — they simply arose from copie s of existing genes. The old version went on doing its job, and the new copy became free to evolve novel functions. Read more
20.05.2014
Tracking colony-building insects

Scientists have long been doing research into how the complicated system of living together in insect colonies functions. An international group of researchers – including scientists from Münster University – have now sequenced and analysed the genome of one type of termite. This means that they have now been able to compare the termites' DNA with that of ants and colony-building bees. The study has been published in "Nature Communications". Read more
09.04.2013
Prestigious award for bioinformatician

Prof. Erich Bornberg-Bauer from the Institute for Evolution and Biodiversity at Münster University has received a prestigious Program Grant from the international "Human Frontier Science Program" (HFSP). With this grant the HFSP supports outstanding scientists from various countries who jointly work on innovative research topics. Read more
12.07.2012
Enigma of Twisted-Wing Parasites resolved

Scientists have long been mystified by the insect group called twisted-wing parasite. It includes over 500 species and although it has been known for almost 200 years, till now it could not be linked to any super-ordinate group of insects. A team of scientists have, for the first time ever, sequenced the entire genetic code – the genome – of a twisted-wing parasite, thus enabling the scientists to classify these insects as a sister group of the beetle. Read more
11.02.2011
Secrets of fungus-growing

An international consortium of researchers, including Prof. Erich Bornberg-Bauer from Münster University's Institute of Evolution and Biodiversity, has been able top demonstrate for the first time that the highly specialized lifestyle of leafcutter ants has been deposited in the insects' genetic make-up. For example, they are missing certain genes which are otherwise necessary for digestion. Read more
18.01.2010
Bacteria change sperm of insects

Es ist ein recht gerissener Trick im Naturreich: Ein Bakterium sorgt dafür, dass sich sein Wirt erfolgreich vermehrt, indem er das Erbgut der Spermien verändert. Darin hatten Forscher überraschenderweise Teile des Bakterien-Genoms gefunden. Mit einem raffinierten genetischen Trick sichert das Bakterium Wolbachia sein Überleben in Insekten: Der Parasit manipuliert die Fortpflanzung seiner Wirte in großem Stil. Auf die Schliche gekommen ist dem Bakterium eine internationale Forschergruppe, die das Erbgut von Erzwespen entziffert und darin ein Gen des Schmarotzers entdeckt hat. Read more